Martin Yanofsky
Research
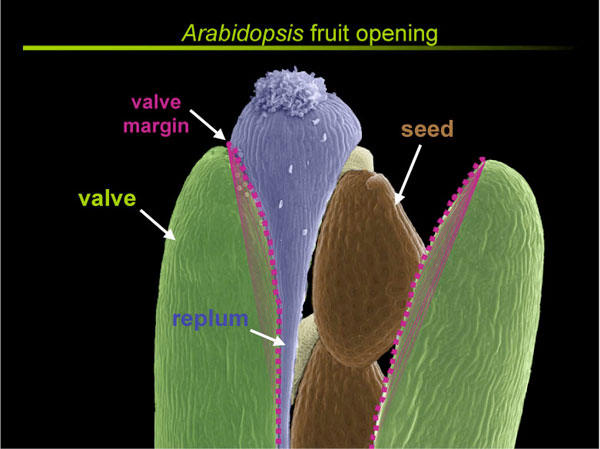
Research in the Yanofsky lab focuses the genes involved in fruit development in Arabidopsis thaliana. The fruit, which is arguably the most complex plant organ, is derived from the fertilized gynoecium (carpels), and consists of a number of distinct tissue types, including the valves, replum and valve margin. The valve margin, which forms at the boundary between the valve and replum regions, consists of a narrow stripe of cells that extends along the entire length of the fruit. Upon fruit maturation, cell separation within the valve margin region allows the valves to detach from the replum and for the seeds to be dispersed. We have shown that the SHATTERPROOF1 ( SHP1) and SHP2 MADS-box genes act redundantly to specify valve margin cell fate. The SHP genes act to promote the expression of the INDEHISCENT ( IND) and ALCATRAZ ( ALC) bHLH genes, which are also required for differentiation of valve margin cells. Because the valve margin fails to differentiate in these mutants, the fruits fail to open and the seeds cannot be dispersed through their normal mechanism. We have also shown that the FRUITFULL ( FUL) MADS-box gene is required for the post-fertilization elongation of the fruit and is required for valve cell differentiation. Interestingly, the FRUITFULL gene product is expressed in valve cells where it negatively regulates expression of SHP, IND, and ALC genes such that the SHP, IND, and ALC genes are ectopically expressed in fruitfull mutant valves. Similarly, the REPLUMLESS ( RPL) gene encodes a homeodomain transcription factor that is required for formation of the replum. RPL is expressed in the replum region where it acts to negatively regulate SHP, IND, and ALC. The combined action of FUL in the valves and RPL in the replum allows for the valve margin identity genes to be expressed in the narrow stripe of cells that differentiate into the valve margin. More recently, we have begun to understand the role of additional fruit development genes leading to a framework model for the genetic interactions that pattern the Arabidopsis fruit.
Our lab has also initiated a project aimed at identifying the major genes that specify the female reproductive tract in Arabidopsis. This region plays a key role in ensuring efficient pollen tube migration as they seek out unfertilized ovules, and yet little is known about the regulatory genes that control the formation of the reproductive tract. We have recently shown that the NO TRANSMITTING TRACT ( NTT) gene encodes a zinc finger transcription factor that is necessary for formation of transmitting tract cells and for preventing the cell death that normally occurs within the transmitting tract region.
Select Publications
- Crawford BC, Sewell J, Golembeski G, Roshan C, Long JA, Yanofsky M.F. (2015) Plant development. Genetic control of distal stem cell fate within root and embryonic meristems. Science. 347:655-9.
- Ripoll, J.J., Bailey, L.J., Mai, Q.A., Wu, S.L., Hon, C.T., Chapman, E.J., Ditta, G.S., Estelle, M., and Yanofsky, M.F. (2015) microRNA regulation of fruit growth. Nature Plants. 1:15036 doi:10.1038/nplants.2015.36.
- Crawford, B.C.W. and Yanofsky, M.F. (2011). HALF FILLED promotes reproductive tract development and fertilization efficiency in Arabidopsis thaliana. Development 138:2999-3009.
- Crawford, B. C. W., Ditta, G., and Yanofsky, M.F. (2007) The NTT gene is required for transmitting-tract development in carpels of Arabidopsis thaliana. Current Biology 17:1101-1108.
- Dinneny, J. R., Weigel, D., and Yanofsky, M.F. (2005) A genetic framework for fruit patterning in Arabidopsis thaliana. Development 132:4687-4696.
- Liljegren, S.J., Roeder, A.H.K., Kempin, S.A., Gremski, K, Østergaard, L., Guimil, S., Khammungkhune, D. and Yanofsky, M.F. (2004) Control of fruit patterning in Arabidopsis by INDEHISCENT. Cell 116:843-853.
Biography
Marty Yanofsky received his Ph.D. from the University of Washington and was an NSF Postdoctoral Fellow in Plant Biology at Caltech. He was the recipient of a Packard Fellowship for Science and Engineering and a Beckman Young Investigator Award. Marty Yanofsky is a member of the National Academy of Sciences.

