Cornelis Murre
Research
We study gene regulation in adult stem cells and developing lymphocytes. Both global and single cell strategies are being utilized with the aim to describe normal development as well as aged and diseased states in molecular terms.
We have a long-standing interest in deciphering the role and regulation of helix-loop-helix proteins in lymphocyte development. This class of factors plays key roles in hematopoiesis. Currently our main interests are in the role of these factors in the control of hematopoietic stem cell homeostasis, B- and T-lineage specification and commitment, aging, inflammatory disease and their roles in the periphery in the response to invading pathogens.
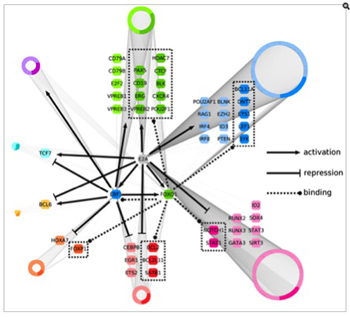
Recent genome-wide studies have identified that a large fraction of the genome transcribed in non-coding regions. We are now faced with the question as to how these large non-coding RNAs relate to the control of gene expression. We have identified a subset of lineage-specific and developmental-stage specific non-coding RNAs. Our interests are to determine the function of these non-coding RNAs and to study their potential roles in modulating long-range chromatin structure and genomic interactions.
Our knowledge of chromatin structure and long-range genomic interactions in mammalian cells is still rudimentary. How do the genomes of lymphocytes, plasma cells and granulocytes adopt such unique and distinct structures? We aim to resolve these questions using genome-wide chromosome-conformation capture studies (HiC) in conjunction with computational approaches to describe the topologies of lymphoid and myeloid genomes in molecular terms.
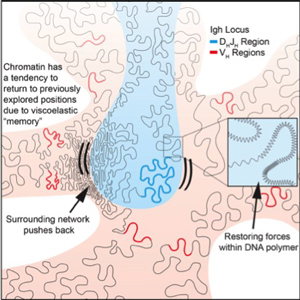
During developmental progression of lymphoid cells coding and regulatory elements interact to induce lineage-specific programs of gene expression. We recently found that in eukaryotic cells coding and regulatory genomic elements bounce back and forth within the chromatin network until specific genomic interactions are established, and that spatial confinement of topological domains largely controls the times for such encounters. Our future studies aim to examine how epigenetic and structural determinants affect the trajectories adopted by the chromatin fiber in living cells and how this relates to genomic encounters involving enhancers and promoters.
We have recently demonstrated that genes encoding for key developmental regulators reposition during developmental progression. We now aim to address the question as to why and how genomic regions encoding for developmental regulator reposition during developmental progression, how their release from the lamina is regulated and how they associate with active transcription factories.
Select Publications
- Isoda, T., Moore, A., He, Chandra, V., Aida, M., Denholtz, van Hamburg, J.P., Fisch, K., Chang, A.N., Fahl, S. Wiest, D.L. and Murre, C. 2017. Non-Coding Transcription Instructs Cohesin-Dependent Chromatin Folding and Compartmentalization to Dictate Enhancer-Promoter Communication and T Cell Fate. Cell 171, 103-119.
- Miyazaki, M., Miyazaki, K., Chen, K., Jin, Y., Turner, J., Moore, A.J., Saito, R., Yoshida, K., Ogawa, S., Rodewald, H.R., Lin, Y.C., Kawamoto, H. and Murre C. 2017. The E-Id protein axis specifies adaptive lymphoid cell identity and suppresses thymic innate lymphoid cell development. Immunity 46, 818-834.
- Zhu, Y., Gong, K., Denholtz, M., Chandra, V., Kamps, M.P., Alber, F. and Murre, C. (2017). Comprehensive characterization of neutrophil genomes. Genes Dev. 31, 141-153.
- Miyazaki. M., Miyazaki, K., Chen, S.Y., Chandra, V., Agata, Y., Chang, A., Kawamoto, H. and Murre, C. 2015. The E-Id protein axis modulates the activities of the PI3K-AKT-mTORC1-Hif1a and c-myc/p19Arf pathways to suppress innate variant TFH cell development, thymocyte expansion and lymphomagenesis. Genes Dev. 29, 409-425.
- Bossen, Murre, C.S., Chang, A.N., Mansson, R., Rodewald, H.R. and Murre, C. 2015. The chromatin remodeler Brg1 activates enhancer repertoires to establish B cell identity and modulate cell growth. Nat. Immunol. 16, 775-784.
- Lucas, J.S., Zhang, J., Dudko, O.S. and Murre, C. 2014. The 3-D trajectories adopted by the immunoglobulin heavy chain locus: First-passage times and long-range genomic interactions. Cell 158, 339-352.
- The 3-D trajectories adopted by coding and regulatory DNA elements: First-passage times for genomic interactions. Lucas JS, Zhang J, Dudko OS, Murre C. Cell 158, 339-352 (2014).
- Id2 and Id3 maintain the regulatory T cell pool to suppress inflammatory disease. Miyazaki M, Miyazaki K, Chen SY, Itoi M, Miller M, Lu L-F, Varki N, Chang A, Broide DH, Murre C. Nat Immunol 15, 767-776 (2014)
- The establishment of B versus T cell identity. Miyazaki K, Miyazaki M, Murre C. Trends Immunol 35, 205-210 (2014)
- Boosting lymphocyte production. Murre C. Immunity 38, 1081-1083 (2013)
- Nuclear location and the control of developmental progression. Lin YC, Murre C. Curr Opion Genet Dev 23, 104-108 (2013)
- A HESitant decision for T cells. Bortnick A, Murre C. Nat Immunol 14, 1209-1210 (2013)
- Global changes in the nuclear positioning of genes and intra- and interdomain genomic interactions that orchestrate B cell fate. Lin YC, Benner C, Mansson R, Heinz S, Miyazaki M, Miyazaki K, Chancera V, Bossen C, Glass CK, Murre C. Nat Immunol 12,1196-1204 (2012)
- Chromatin topology and the regulation of antigen receptor assembly. Bossen C, Mansson R, Murre C . Annual Rev Immunol 30, 337-356 (2012)
- Positive intergenic feedback circuitry, involving EBF1 and FOXO1, orchestrates B-cell fate. Mansson R, Welinder E, Ashber J, Lin YC, Benner C, Glass CK, Sigvardsson M, Murre C. Proc Natl Acad. Sci USA 108, 17402-17407 (2012)
- Multilineage priming of enhancer repertoires precedes commitment to the B and myeloid cell lineage in hematopoietic progenitors. Mercer EM, Lin YC, Jhunjhunwala S, Dutkowski J, Flores M, Sigvardsson M, Ideker T, Glass CK, Murre C. Immunity 35, 413-425 (2011)
- The opposing roles of the transcription factor E2A and its antagonist Id3 that orchestrate and enforce the naïve fate of T cells. Miyazaki M, Rivera RR, Miyazaki K, Lin YC, Agata Y, Murre C. Nat Immunol 12, 992-1001 (2011)
- E2A and HEB act in concert to induce the expression of FOXO1 in the common lymphoid progenitor . Welinder E, Mansson R, Mercer EM, Bryder D, Sigvardsson M, Murre C. Proc Natl Acad Sci USA 108, 17402-17407 (2011)
- CTCF-binding elements mediate control of V(D)J recombination. Guo C, Yoon HS, Franklin A, Jain S, Ebert A, Cheng HL, Hansen E, Despo O, Bossen C, Vettermann C, Bates JG, Richards N, Myers D, Patel H, Gallagher M, Schlissel MS, Murre. C, Busslinger M, Giallourakis CC, Alt FW. Nature 477, 424-430 (2011)
- Transcription and recombination factories: Common features? . Lucas JL, Bossen C, Murre C. Curr Opin Cell Biol 23, 318-324 (2011)
- Ldb1, a new guardian of hematopoietic stem cell maintenance. Welinder E, Murre C. Nat Immunol 12, 113-114 (2011)
- CCCTC-binding factor (CTCF) and cohesin influence the genomic architecture of the Igh locus and antisense transcription in pro-B cells. Degner SC, Verma-Gaur J, Wong TP, Bossen C, Iverson GM, Torkamani A, Vettermann C, Lin YC, Ju Z, Schulz D, Murre CS, Birshtein BK, Schork NJ, Schlissel MS, Riblet R,. Murre C, Feeney AJ Proc Natl Acad Sci USA 108, 9566-9571 (2011)
- Factors and networks that underpin early hematopoiesis. Mercer EM, Lin YC, Murre C. Semin Immunol 5, 317-325 (2011)
- A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Lin YC, Jhunjhunwala S, Benner C, Heinz S, Welinder E, Mansson R, Sigvardsson M, Hagman J, Espinoza A, Dutkowski J, Ideker T, Glass CK, Murre C. Nat Immunol 11, 635-643 (2010)
- Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK. Mol Cell 38, 576-89 (2010)
- Chromatin architecture and the generation of antigen receptor diversity. Jhunjhunwala S, van Zelm MC, Peak M, Murre C. Cell 138, 435-448 (2009)
- Distinct roles for E12 and E47 in B cell specification and the sequential rearrangement of immunoglobulin light chain loci. Beck K, Peak M, Ota T, Nemazee D. Murre C. J Exp Med 206, 2271-2284 (2009)
- E2A proteins maintain the hematopoietic stem cell pool and promote the maturation of myelolymphoid and myeloerythroid progenitors. Semerad CL, Mercer EM, Inlay MA, Weissman IL, Murre C. Proc Natl Acad Sci USA 106, 1930-1935 (2009)
- Developmental trajectories in early hematopoiesis. Murre C . Genes Devel 15, 2366-2370 (2009)
- The 3D-structure of the immunoglobulin heavy chain locus: Implications for long-range genomic interactions. Jhunjhunwala S, van Zelm MC, Peak M, Cutchin S, Riblet R, van Dongen JJM, Grosveld F, Knoch TA, Murre C. Cell 133, 265-279 (2008)
Biography
Kees Murre performed his graduate work at Harvard Medical School and was a postdoctoral fellow at MIT. He is a Searle Scholar and the recipient of the National Institutes of Health Merit Award.

