Douglass Forbes
Research
Our lab has two primary areas of interest: (1) the structure, function and assembly of the vertebrate nucleus, and (2) the mechanism by which the cell regulates where and how to assemble major mitotic structures, such as the mitotic spindle, the nuclear membranes and nuclear pore complexes.
The Nucleus: Structure, Function and Assembly
A focus of the research in my laboratory is the eukaryotic nucleus: its structure and function. We work both in vivo and in vitro. For in vivo studies, we use mammalian cultured cell lines to examine nuclear transport or assembly using RNAi technology to knock down expression of genes of interest to determine their effect on function. For in vitro studies, we use a nuclear reconstitution system which efficiently assembles nuclei in the test tube from soluble and membrane components. This system makes use of an extract of Xenopus eggs, which naturally store abundant amounts of disassembled nuclear components in preparation for the rapid cleavages of early division. Upon addition of DNA or chromatin to the in vitro interphase extract, nuclei containing nuclear membranes, a nuclear lamina, and nuclear pores quickly assemble. The in vitro reconstituted nuclei are capable of nuclear import, DNA replication, pol III transcription, and even mitotic disassembly. In parallel, we use a mitotic egg extract to study spindle and kinetochore assembly.
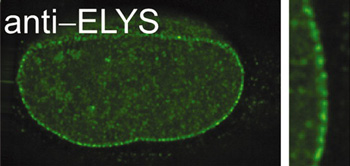
A long-standing interest of the laboratory has been the nuclear pore, a large macromolecular complex of 120 million daltons that spans the nuclear membranes. All communication between the nucleus and cytoplasm occurs through the nuclear pore. In vivo, the pore complex imports nuclear proteins and exports mRNA, tRNA, snRNAs, and ribosomal precursors. We study: (1) assembly of the nucleus itself, (2) assembly of nuclear pores, (3) the regulation of these processes, and (4) the mechanism of nuclear transport, including mRNA export, protein export, and protein import. Using antibodies we can immunodeplete a nuclear assembly extract of individual nuclear pore proteins, then reconstitute nuclei which lack that protein. Such "designer" nuclei can be tested for alterations in nuclear pore structure, pore function, or pore assembly. In this way we discovered the protein responsible for initiating pore assembly at the nucleus (Rasala et al, 2006, 2008). This protein, ELYS, acts by binding to chromatin at AT-rich sequences and recruiting key structural subunits of the nuclear pore to these sites to initiate nuclear pore assembly. With experiments of this type, we hope to molecularly dissect and elucidate the gates that "guard the fortress" of the genome.
Regulation of the Major Mitotic Assembly Events: Cellular GPSs
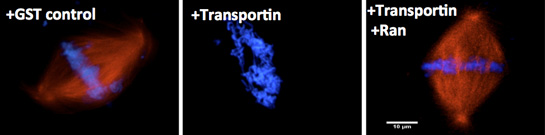
The second major interest of the laboratory involves understanding the regulation of assembly of the major mitotic structures. Specifically, we are focused on how assembly is regulated such that the mitotic spindle, and later the nuclear membranes and nuclear pores, form only around chromatin, i.e., in a spatially precise manner. We have shown that an abundant cellular protein, Importin beta, acts in mitosis to negatively regulate both nuclear membrane fusion and nuclear pore assembly everywhere except in the vicinity of the chromosomes (Harel et al, M.B.C, 2003). The small GTPase Ran, produced in its GTP-bound form only near chromatin, acts as the counteracting positive regulator, a “GPS” to tell where assembly should occur. We have extended this work to Transportin, a second karyopherin. We believe that Importin beta and Transportin act as global regulators of cellular events involving the genome, including those controlling nuclear import, spindle formation, correct nuclear membrane assembly, and nuclear pore assembly.
Select Publications
- Forbes, D.J., Travesa, A., Nord, M., and C. Bernis. (2015). Nuclear transport factors: Global regulation of mitosis. Current Opinion in Cell Biology 35:78-90.
- Schwartz, M., Travesa, A., Martell, S., and D.J. Forbes. (2015). Analysis of the initiation of nuclear pore assembly by ectopically targeting nucleoporins to chromatin. Nucleus 6:40-54
- Bernis, C., Swift-Taylor, B., Nord, M., Carmona, S., Chook, Y.M., and Forbes, D.J. (2014). Transportin acts to regulate mitotic assembly events by target binding rather than Ran sequestration. Molecular Biology of the Cell 25: Jan 29. [Epub ahead of print].
- Bernis, C. and Forbes, D.J. (2014). Analysis of nuclear reconstitution, nuclear envelope assembly, and nuclear pore assembly using Xenopus in vitro assays. In: “Nuclear Pore Complex and Nucleocytoplasmic Transport”, Methods in Cell Biology, Volume 122, Chapter 8, pp. 165-191. Ed. Valérie Doye, Elsevier Press (in press)
- Powers, M. and Forbes, D.J. (2013). Nucleocytoplasmic transport: Beginning to gel? Current Biology 22: R1006-1009.
- Fichtman, B., Ramos, C., Rasala, B., Harel, A., and Forbes, D.J. (2010). Inner/outer nuclear membrane fusion in nuclear pore assembly: Biochemical demonstration and molecular analysis. Molecular Biology of the Cell 21:4197-4211. Lachish-Zalait, A., Lau, C., Fichtman, B., Zimmerman, E., Harel, A., Gaylord, M., Forbes, D.J, and Elbaum, M. (2009). Transportin mediates nuclear entry of DNA in vertebrate systems. Traffic 10:1414-1428
- Lau, C.K., Delmar, V.A., Chan, R.C., Phung, Q., Bernis, C., Fichtman, B., Rasala, B.A., Forbes. D.J. (2009). Transportin regulates major mitotic assembly events: From spindle to nuclear pore assembly. Molecular Biology of the Cell 20:4043-4058
- Brown, C., Kennedy, C., Delmar, V., Forbes, D.J. and P. Silver. (2008). Global histone acetylation induces functional genomic reorganization at mammalian nuclear pores. Genes and Development 22:627-639.
- Chakraborty, P., Wang, Y., Wei, J-H. , van Deursen, J., Yu, H., Malureanu, L., Dasso, M., Forbes, D.J., Levy, D., Seemann, J. , and B. Fontoura. (2008). Nucleoporin levels regulate cell cycle progression and phase-specific gene expression. Developmental Cell 15:657-667.
- Rasala, B., Ramos, C., Harel, A., and Forbes, D.J. (2008). Capture of AT-rich chromatin by ELYS recruits POM121 and NDC1 to initiate nuclear pore assembly. Molecular Biology of the Cell 19:3982-3996
- Orjalo, A., Arnaoutov, A. , Shen, Z., Boyarchuk, Y., Zeitlin, S., Fontoura, B., Briggs, S., Dasso, M., and Forbes, D.J. (2006). The Nup107-160 nucleoporin complex is required for correct bipolar spindle assembly. Molecular Biology of the Cell 17: 3806-3818.
- Rasala, B., Orjalo, A., Shen, Z., Briggs, S., and Forbes, D.J. (2006). ELYS is a dual nucleoporin/kinetochore protein required for nuclear pore assembly and proper cell division. Proc. Natl. Acad. Sci., U.S.A. 103:17801-17806.
- Harel, A. and Forbes, D.J. (2004). Importin beta: Conducting a much larger cellular symphony. Molecular Cell 16:319-330.
- Harel, A., Chan, R., Lachish-Zalait, A., Zimmerman, A., Elbaum, M., and Forbes, D.J. (2003). Importin beta negatively regulates nuclear membrane fusion and NPC assembly. Molecular Biology of the Cell 14:4387-4396.
- Harel, A., Orjalo, A., Vincent, T., Lachish-Zalait, A., Vasu, S., Shah, S., Zimmerman, A., Elbaum, M. and Forbes, D.J. (2003). Removal of a single pore subcomplex results in vertebrate nuclei devoid of nuclear pores. Molecular Cell 11:853-864.
Biography
Douglass Forbes received her Ph.D. from the University of Oregon and was an American Cancer Society postdoctoral fellow at UCSF in the Dept. of Biochemistry and Biophysics. She was a PEW Fellow in the Biomedical Sciences, and an elected member of the Governing Council of the American Society of Cell Biology. Dr. Forbes served as the Vice Chair of Cell and Developmental Biology from 2000-2005 and 2007-2009. She has served on multiple editorial boards, both past and present.

