Deborah Yelon
Research
Organogenesis requires the carefully orchestrated arrangement of multiple cell types into a specific three-dimensional form that is essential for effective organ function. The embryonic zebrafish heart provides a clear example of the relationship between organ form and function: it is composed of two major chambers, a ventricle and an atrium, each with discrete morphological and physiological characteristics that define its functional capacity. The goal of our laboratory's research is to understand the molecular mechanisms that specify the size, shape, and attributes of each cardiac chamber. By taking advantage of the unique arsenal of genetic and embryologic experimental approaches available in zebrafish, we can identify crucial regulators of chamber formation and determine their precise impacts on cell fate and cell behavior. In the long term, this work will improve our comprehension of the etiology of congenital heart disease and inform strategies for regenerative medicine and tissue engineering.
Regulation of heart size
Specification of the proper number of cardiac progenitor cells is a critical determinant of heart size, but we do not fully understand the network of signals that regulate the dimensions of the cardiac progenitor pool. Several types of secreted factors, including BMP, FGF, and Hedgehog, are implicated in promoting the assignment of cardiac identity. However, much less is known about the opposing factors that set limits for cardiac progenitor specification. We are therefore particularly interested in zebrafish mutants with large hearts composed of too many cardiomyocytes. Notably, our recent analyses of these mutants have revealed two potent mechanisms for restricting the formation of cardiac progenitors. These studies indicate that generation of the proper number of progenitors involves interplay between inductive and repressive pathways.
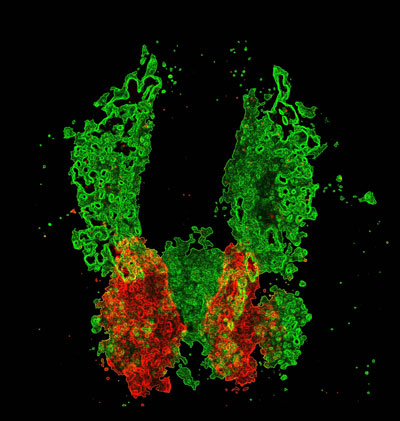
Cardiac cells (red) develop in proximity to the vascular plexi (green) prior to heart formation. Allocation of cardiac and vascular progenitors requires interplay between cardiovascular pathways: components of vessel and blood specification pathways repress formation of cardiac progenitors and thereby limit heart size (Schoenebeck et al., 2007).
Regulation of heart shape
An elaborate series of cell movements and cell shape changes creates the characteristic contours of the heart, and all of these cellular activities can be influenced by the surrounding environment. However, it is not clear which extrinsic and intrinsic cues regulate cell behaviors during chamber morphogenesis. By combining high-resolution live imaging with genetic analysis, we can elucidate pathways with a crucial influence on the actions of individual cardiomyocytes. To date, we have focused our attention on two particular steps of cardiac morphogenesis: the midline merger of the bilateral cardiomyocyte precursor populations and the emergence of chamber curvatures. During both of these processes, regionally restricted patterns of cell behavior underlie key features of cardiac morphology, and interactions of the cardiomyocytes with their environment have a significant impact on morphogenetic control.
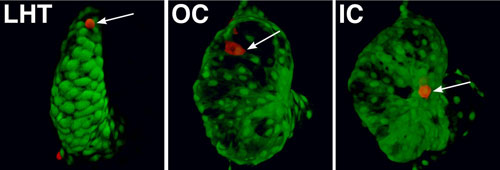
Regionally confined cell shape changes underlie the emergence of chamber curvatures. A portion of the linear heart tube (LHT) transforms into an expanded ventricle with a bulging outer curvature (OC) and a recessed inner curvature (IC). Individual cells (red, arrows) are small and round in the LHT. Cells in the OC become flattened and elongated, whereas cells in the IC remain relatively cuboidal, creating the characteristic shape of the ventricle (Auman et al., 2007).
Select Publications
- Keegan, B.R., J. Feldman, G. Begemann, P. Ingham, and D. Yelon. (2005) Retinoic acid signaling restricts the cardiac progenitor pool. Science 307: 247-249.
- Auman, H.J., H. Coleman, H.E. Riley, F. Olale, H.-J. Tsai, and D. Yelon. (2007) Functional modulation of cardiac form through regionally confined cell shape changes. PLoS Biology 5: e53.
- Schoenebeck, J.J., B.R. Keegan, and D. Yelon. (2007) Vessel and blood specification override cardiac potential in anterior mesoderm. Developmental Cell 13: 254-267.
- Waxman, J.S., B.R. Keegan, R.W. Roberts, K.D. Poss, and D. Yelon. (2008) Hoxb5b acts downstream of retinoic acid signaling in the forelimb field to restrict heart field potential in zebrafish. Developmental Cell 15: 923-934.
- Garavito-Aguilar, Z.V., H.E. Riley, and D. Yelon. (2010) Hand2 ensures an appropriate environment for cardiac fusion by limiting fibronectin function. Development 137: 3215-3220.
- Totong, R., T. Schell, F. Lescroart, L. Ryckebüsch, Y.-F. Lin, T. Zygmunt, L. Herwig, A. Krudewig, D. Gershoony, H.-G. Belting, M. Affolter, J. Torres-Vázquez, and D. Yelon. (2011) The novel transmembrane protein Tmem2 is essential for coordination of myocardial and endocardial morphogenesis. Development 138: 4199-4205
- Lin, Y.-F., I. Swinburne, and D. Yelon. (2012) Multiple influences of blood flow on cardiomyocyte hypertrophy in the embryonic zebrafish heart. Developmental Biology 362:242-253.
- Targoff, K.L., S. Colombo, V. George, T. Schell, S.-H. Kim, L. Solnica-Krezel, and D. Yelon. (2013) Nkx genes are essential for maintenance of ventricular identity. Development 140:4203-4213.
- Schumacher, J.A., J. Bloomekatz, Z.V. Garavito-Aguilar, and D. Yelon. (2013) tal1 regulates the formation of intercellular junctions and the maintenance of identity in the endocardium. Developmental Biology 383:214-226.
- Zeng, X.-X.I. and D. Yelon. (2014) Cadm4 restricts the production of cardiac outflow tract progenitor cells. Cell Reports 7:951-960.
- Schindler, Y.L., K.M. Garske, J. Wang, B.A. Firulli, A.B. Firulli, K.D. Poss, and D. Yelon. (2014) Hand2 elevates cardiomyocyte production during zebrafish heart development and regeneration. Development 141:3112-3122.
Biography
Deborah Yelon received her Ph.D. in 1996 from Harvard University, where she studied T cell development with Leslie Berg. She then began to investigate heart development in zebrafish as a Life Sciences Research Foundation postdoctoral fellow with Didier Stainier at the University of California, San Francisco. She was a member of the faculty of the Skirball Institute of Biomolecular Medicine at New York University School of Medicine from 2000 to 2009. She has received a Burroughs Wellcome Fund Career Award and an American Heart Association Established Investigator Award.

