Julian Schroeder
Research
Identifying the basic molecular mechanisms by which plants respond to and mount resistance to environmental stresses is fundamental to understanding stress resistance mechanisms to these “abiotic” stresses in plants and is an important goal for developing future strategies for engineering stress resistance in plants. Several abiotic stress mechanisms that we are characterizing are directly linked to water, including drought stress-induced signal transduction mechanisms, salinity resistance mechanisms and how plants respond to heat waves and the continuing rise in the atmospheric CO2 concentration.
Our laboratory’s research is directed at the signal transduction mechanisms and pathways that mediate resistance to environmental (“abiotic”) stresses in plants, including elevated CO2, drought, salinity stress, heat waves and heavy metal stress. These abiotic stresses have substantial negative impacts and reduce plant growth and agricultural productivity. Resistance or improved performance in response to these environmental stresses is relevant for meeting the food and energy needs of the growing human population.
Our research is elucidating the molecular and cell biological stress-induced signal transduction cascades in higher plant cells, examining the chain of events by which plant cells respond to elevated CO2, the drought stress hormone abscisic acid, elevated temperatures and salinity stress to mount specific resistance and adaptation responses. We have developed and adapted interdisciplinary and systems biological approaches to guard cells, which control water loss (transpiration) and CO2 intake in plants and which have become a key model system for understanding dynamic cellular signal transduction and ion channel functions in plants.
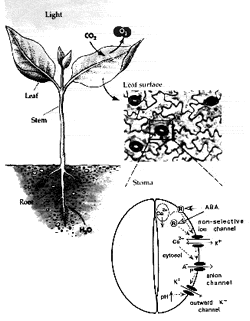
Stomatal pores in the epidermis of leaves allow CO2 influx into leaves from the atmosphere and also mediate transpirational water loss of plants (see figure). Two guard cells surround each pore and control the opening and closing of stomata. In guard cells, cell biological, molecular, patch clamp, epigenetic and time-resolved calcium, abscisic acid and protein kinase activity imaging studies on genetic signaling mutants in Arabidopsis are allowing us to identify and characterize stress-induced signal transduction mechanisms and cascades. We are combining these analyses with new genomic, systems, bioinformatic and proteomic approaches towards discovering new signaling mechanisms and principles. We have identified unique stomatal CO2 sensors and early CO2 signal transduction mechanisms, including ion channels in guard cells through which elevated CO2 reduced stomatal pore apertures. We have identified new early signal transduction mechanisms and contributed to the characterization and co-identification of receptors for the plant stress hormone abscisic acid and have obtained molecular genetic, cell biological, genomic, biophysical whole plant physiological evidence for new genes and mechanisms in guard cells that reduce water loss of Arabidopsis during drought.
A second effort in the lab focuses on identifying genes that mediate salt (sodium) stress resistance and heavy metal uptake and detoxification in plants. In this research we identified the plant HKT transporter family and showed its central role in mediating salinity resistance in the reference plant, Arabidopsis thaliana, by limiting sodium transfer to leaves via xylem removal. Research on the staple crops rice and wheat and other plant species is showing that this same HKT transporter mechanism plays a major role in determining salinity resistance. HKT gene-focused molecular breeding efforts are indicating major improvements in yield, illustrating how basic Arabidopsis research is leading to innovation in agriculture.
Our research into heavy metal stress led to the parallel discovery of the genes encoding the central heavy metal detoxification enzymes in plants, phytochelatin synthases. Furthermore, the long sought family of transporters that mediate heavy metal accumulation in plant vacuoles were identified in collaboration with experts. These basic research advances can provide key tools for avoiding toxic heavy metal and arsenic accumulation in edible plant tissues, a problem facing millions of people today leading to cancer and other diseases. Furthermore, these basic research advances can contribute key tools for engineering plants for environmental remediation (bioremediation) by removal of heavy metals from soils.
Members in our lab are being trained in interdisciplinary and systems biological techniques while pursuing individual research projects.
Go to full publication list
Select Publications
- Pankasem N, Hsu PK, Lopez BNK, Franks PJ, Schroeder JI. Warming triggers stomatal opening by enhancement of photosynthesis and ensuing guard cell CO2 sensing, whereas higher temperatures induce a photosynthesis-uncoupled response. New Phytol. Dec;244(5):1847-1863. doi: 10.1111/nph.20121. (2024)
- Seller CA, Schroeder JI. Distinct guard cell-specific remodeling of chromatin accessibility during abscisic acid- and CO2-dependent stomatal regulation. Proc Natl Acad Sci U S A. Dec 26;120(52):e2310670120. doi: 10.1073/pnas.2310670120. (2023)
- Takahashi Y, Bosmans KC, Hsu PK, Paul K, Seitz C, Yeh CY, Wang YS, Yarmolinsky D, Sierla M, Vahisalu T, McCammon JA, Kangasjärvi J, Zhang L, Kollist H, Trac T, Schroeder JI. Stomatal CO2/bicarbonate sensor consists of two interacting protein kinases, Raf-like HT1 and non-kinase-activity activity requiring MPK12/MPK4. Science Adv. Dec 9;8(49):eabq6161. doi: 10.1126/sciadv.abq6161. (2022)
- Waadt R., Seller, C.A., Hsu, P-K., Takahashi Y., Munemasa S., Schroeder J.I., Plant hormone regulation of abiotic stress responses. Nature Reviews Mol. Cell Biol. doi: 10.1038/s41580-022-00479-6. PMID 35513717. (2022)
- Hsu PK, Takahashi Y, Merilo E, Costa A, Zhang L, Kernig K, Lee KH, Schroeder JI. Raf-like kinases and receptor-like (pseudo)kinase GHR1 are required for stomatal vapor pressure difference response. Proc Natl Acad Sci U S A. 118(47):e2107280118. doi: 10.1073/pnas.2107280118. (2021) PMID: 34799443
- Zhang*, L., Takahashi*, Y., Hsu, P. K., Kollist, H., Merilo, E., Krysan, P. J., & Schroeder, J. I. FRET kinase sensor development reveals SnRK2/OST1 activation by ABA but not by Me-Jasmonate and high CO2 during stomatal closure. (*equally contributing authors) eLife 2020;9:e56351. doi:10.7554/eLife.56351. (2020). PMC7289597
- Takahashi, Y., Zhang, J., Hsu, P.K., Ceciliato, P.O.H., Zhang, L., Dubeaux, G., Munemasa, S., Ge, C, Zhao, Y., Hauser, F., & Schroeder, J.I. MAP3Kinase-dependent SnRK2-kinase activation is required for abscisic acid signal transduction and rapid osmotic stress response. Nature Communications Jan 2;11(1):12. doi: 10.1038/s41467-019-13875-y. (2020). PMC6940395
- Bailey-Serres, J., Parker, J. E., Ainsworth, E. A., Oldroyd, G. E. D., Schroeder, J. I., Genetic strategies for improving crop yields. Nature 575 (7781): 109-118. doi: 10.1038/s41586-019-1679-0. (2019). PMC7024682
- Hauser* F, Ceciliato* PHO, Lin YC, Guo D, Gregerson JD, Abbasi N, Youhanna D, Park J, Dubeaux G, Shani E, Poomchongkho N, Schroeder J I. A seed resource for screening functionally redundant genes and isolation of new mutants impaired in CO2 and ABA responses. .(*equally contributing authors) J Exp Bot 70(2):641-651. doi: 10.1093/jxb/ery363. (2019) PMC6322574
- Zhang J, Wang N, Miao Y, Hauser F, McCammon JA, Rappel WJ, Schroeder JI. Identification of SLAC1 anion channel residues required for CO2/bicarbonate sensing and regulation of stomatal movements. Proc Natl Acad Sci U S A. 115(44):11129-11137 doi: 10.1073/pnas.1807624115. (2018). PMC6217375
- Schroeder, J. I. (6 individual author commentaries by J. Dangl, A.Osbourn et al.) What is the Next Frontier in Plant Engineering? Coping with High CO2. Cell 74(3):499-500 doi: 10.1016/j.cell.2018.07.012. (2018). PMC6541440
- Hauser F, Li Z, Waadt R, Schroeder JI. SnapShot: Abscisic Acid Signaling. Cell 171(7): 1708-1708 doi: 10.1016/j.cell.2017.11.045. PMID: 29245015 (2017).
- Li Z, Waadt R, Schroeder JI. Release of GTP Exchange Factor Mediated Down-Regulation of Abscisic Acid Signal Transduction through ABA-Induced Rapid Degradation of RopGEFs. PLoS Biol. May 18;14(5):e1002461. doi: 10.1371/journal.pbio.1002461. eCollection (2016).
- Stephan A. B., Kunz H. H., Yang E. & Schroeder J.I. Rapid hyperosmotic-induced Ca2+ responses in Arabidopsis thaliana exhibit sensory potentiation and involvement of plastidial KEA transporters. Proc Natl Acad Sci USA pii: 201519555 (2016).
- Wang C, Hu H, Qin X, Zeise B, Xu D, Rappel WJ, Boron WF, Schroeder JI. Reconstitution of CO2 regulation of SLAC1 anion channel and function of CO2-permeable PIP2;1 aquaporin as carbonic anhydrase 4 interactor. Plant Cell. (2016).
- Brandt B, Munemasa S, Wang C, Nguyen D, Yong T, Yang PG, Poretsky E, Belknap TF, Waadt R, Alemán F, Schroeder JI. Calcium specificity signaling mechanisms in abscisic acid signal transduction in Arabidopsis guard cells. Elife. Jul 29;4. doi: 10.7554/eLife.10328. (2015).
- Engineer, C. B., Ghassemian, M., Anderson, J., Hu, H., Peck, S. & Schroeder, J. I. Carbonic anhydrases, EPF2 and a novel protease mediate CO2 control of stomatal development. Nature 513: 246-250 (2014).
- Waadt, R., Hitomi, K., Nishimura, N., Hitomi, C., Adams, S. R., Getzoff, E. D. & Schroeder, J. I. FRET-based reporters for the direct visualization of abscisic acid concentration changes and distribution in Arabidopsis. eLife 3:e01739 doi: 10.7554/eLife.01739 (2014).
- Kunz, H. H., Gierth, M., Herdean, A., Cruz, M., Kramer, D.M., Spetea, C. & Schroeder, J. I. Plastidial Transporters KEA1, 2 and 3 are essential for chloroplast osmoregulation, integrity and pH regulation in Arabidopsis. Proc Natl Acad Sci USA 111: 7480-7485 (2014).
- Jones, A.M., Xuan, Y., Xu, M., Wang, R.-S., Ho, C.-H., Lalonde, S., You, C.H., Sardi, M.I., Parsa, S.A., Smith-Valle, E., Su, T., Frazer, K.A., Pilot, G., Pratelli, R., Grossmann, G., Acharya, B.R., Hu, H.-C., Engineer, C., Villiers, F., Ju, C., Takeda, K., Su, Z., Dong, Q., Assmann, S.M., Chen, J., Kwak, J.M., Schroeder, J.I., Albert, P., Rhee, S.Y., Frommer, W.B. Border control – a membrane-linked interactome of Arabidopsis. Science 344: 711-716. (2014).
- Hauser F, Chen W, Deinlein U, Chang K, Ossowski S, Fitz J, Hannon GJ, Schroeder JI. A Genomic-Scale Artificial MicroRNA Library as a Tool to Investigate the Functionally Redundant Gene Space in Arabidopsis. Plant Cell 25: 2848-2863 (2013).
- Schroeder, J.I., Delhaize, E., Frommer, W.B., Guerinot, M.L., Harrison, M.J., Herrera-Estrella,L., Horie, T., Kochian, L.V., Munns, R., Nishizawa, N.K., Tsay, Y.F. & Sanders, D. Using Membrane Transporters to Improve Crops for Sustainable Food Production. Nature 497: 60-66 (2013).
- Brandt, B., Brodsky, D.E., Xue, S., Negi, J., Iba, K., Kangasjärvi, J., Ghassemian, M., Stephan, A.B., Hu, H., Schroeder, J.I. Reconstitution of abscisic acid activation of SLAC1 anion channel by CPK6 and OST1 kinases and branched ABI1 PP2C phosphatase action. Proc Natl Acad Sci USA 109:10593-10598 (2012).
- Kim, T.H., Kunz, H.H., Bhattacharjee, S., Hauser, F., Park, J., Engineer, C., Liu, A., Ha, T., Parker, J.E., Gassmann, W., Schroeder, J.I. Natural Variation in Small Molecule-Induced TIR-NB-LRR Signaling Induces Root Growth Arrest via EDS1- and PAD4-Complexed R Protein VICTR in Arabidopsis. Plant Cell 24: 5177-5192 (2012).
- Kim, T.H., Hauser, F., Ha T., Xue S., Böhmer M., Nishimura N., Munemasa S., Hubbard K., Peine N., Lee B.H., Lee S., Robert N., Parker J.E., Schroeder J.I. Chemical genetics reveals negative regulation of abscisic Acid signaling by a plant immune response pathway. Current Biology 21: 990-997 (2011).
- Xue, S., Hu, H., Ries, A., Merilo, E., Kollist, H. & Schroeder, J.I. Central functions of bicarbonate in S-type anion channel activation and OST1 protein kinase in CO2 signal transduction in guard cell. EMBO J. 30: 1645-1658 (2011).
- Hauser, F., Waadt, R & Schroeder, J.I. Evolution of abscisic acid synthesis and signaling mechanisms. Current Biology 21: R346-355 (2011).
- Song, W.Y., Park, J., Mendoza-Cózatl, D.G., Suter-Grotemeyer, M., Shim, D., Hörtensteiner, S., Geisler, M., Weder, B., Rea, P.A., Rentsch, D., Schroeder, J.I., Lee, Y., Martinoia, E. Arsenic tolerance in Arabidopsis is mediated by two ABCC-type phytochelatin transporters. Proc Natl Acad Sci USA 107:21187-21192 (2010).
- Hubbard, K., Nishimura, N., Hitomi, K., Getzoff, E.D. and Schroeder, J.I. Early Abscisic Acid Signal Transduction Mechanisms. Genes & Development 24: 1695-1708 (2010).
- Hu., H., Boisson-Dernier, A., Israelsson -Nordström, M., Böhmer, M., Xue, S., Ries, A., Godoski, J., Kuhn, J.M. & Schroeder, J.I. Carbonic Anhydrases are Upstream Regulators of CO2-controlled Stomatal Movements in Guard Cells. Nature Cell Biol. 12: 87-93 (2010).
- Mendoza-Cozatl, D.G., Zhai, Z., Jobe, T.O., Akmakjian, G.Z., Song, W.Y., Limbo, O., Russell, M.R., Kozlovskyy, V.I., Martinoia, E., Vatamaniuk, O.K., Russell, P., Schroeder, J.I. Tonoplast-localized Abc2 Transporter Mediates Phytochelatin Accumulation in Vacuoles and Confers Cadmium Tolerance. J. Biol. Chem. (2010).
- Kim, T.H., Böhmer, M., Hu, H., Nishimura, N. & Schroeder, J.I. Guard cell signal transduction network; advances in understanding abscisic acid, CO2, and Ca2+ signaling Annual Reviews of Plant Biol. 61: 561-91 (2010).
- Nishimura, N., Hitomi K, Arvai A.S., Rambo R.P., Hitomi C., Cutler S.R., Schroeder J.I., Getzoff E.D. Structural mechanism of abscisic acid binding and signaling by dimeric PYR1. Science 326:1356-7 (2009).
- Nishimura N., Sarkeshik A., Nito K., Park S.Y., Wang A., Carvalho P.C., Lee S., Caddell D.F., Cutler S.R., Chory J., Yates J.R., Schroeder J.I. PYR/PYL/RCAR family members identified as major in-vivo ABI1 protein phosphatase 2C-interacting proteins in Arabidopsis. Plant Journal 61: 290-299 (2010).
- Park, S.Y., Fung, P., Nishimura, N., Jensen, D.R., Fujii, H., Zhao, Y., Lumba, S., Santiago, J., Rodrigues, A., Chow, T.F., Alfred, S.E., Bonetta, D., Finkelstein, R., Provart, N.J., Desveaux, D., Rodriguez, P.L., McCourt, P., Zhu, J.K., Schroeder, J.I., Volkman, B.F., Cutler, S.R. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324: 1068-1071 (2009).
- Siegel, R.S., Xue, S., Murata, Y., Yang, Y., Nishimura, N., Wang, A. & Schroeder, J.I. Calcium elevation-dependent and attenuated resting calcium-dependent abscisic acid induction of stomatal closure and abscisic acid-induced enhancement of calcium sensitivities of S-type anion and inward-rectifying K channels in Arabidopsis guard cells. Plant Journal 59(2): 207-20 (2009).
- Ward, J.M., Mäser, P., Schroeder, J.I. Plant Ion Channels: Gene Families, Physiology, and Functional Genomics Analyses. Annual Reviews of Physiology 71 (2009).
- Vahisalu, T., Kollist, H., Wang, Y.-F., Nishimura, N., Chan, W.-Y., Valerio, G., Lamminmaki, A., Brosche, M., Moldau, H., Desikan, R., Schroeder, J.I., Kangasjarvi, J. SLAC1 is required for plant guard cell S-type anion channel function in stomatal signalling Nature 452:487-491 (2008).
- Boisson-Dernier, A., Frietsch, S., Kim, T.H., Dizon, M.B. and Schroeder, J.I. The peroxin ABSTINENCE BY MUTUAL CONSENT disrupts male-female gametophyte recognition. Current Biology 18:63-68 (2008).
- Imaizumi, T., Kay, S., Schroeder, J.I. Daily watch on Metabolism. Science 318:1730-1731 (2007).
- Horie, T., Costa, A., Kim, TH, Han, MH, Horie, R., Leung, HY, Miyao, A., Hirochika, H., An, G. and Schroeder, J.I. Rice Os HKT2;1 transporter mediates large Na+ influx component into K+-starved roots for growth. EMBO J. 26:3003-3014 (2007).
- Suh, S.J., Wang, Y.F., Frelet, A., Leonhardt, N., Klein, M., Forestier, C., Mueller-Roeber, B., Cho, M.H., Martinoia, E., Schroeder, J.I. The ATP Binding Cassette Transporter AtMRP5 Modulates Anion and Calcium Channel Activities in Arabidopsis Guard Cells. J Biol Chem 282:1916-1924 (2007).
- Mori, I.C., Murata, Y., Yang, Y., Munemasa, S., Wang, Y.F., Andreoli, S., Tiriac, H., Alonso, J.M., Harper, J.F., Ecker, J.R., Kwak, J.M., Schroeder, J.I. (2006). CDPKs CPK6 and CPK3 Function in ABA Regulation of Guard Cell S-Type Anion- and Ca(2+)- Permeable Channels and Stomatal Closure. PloS Biol. 4.
- Schroeder, J.I., Nambara, E. (2006). A quick release mechanism for abscisic acid. Cell 126:1023-1025.
- Young, J.J., Mehta, S., Israelsson, M., Godoski, J., Grill, E., Schroeder, J.I. (2006). CO2 signaling in guard cells: Calcium sensitivity response modulation, a Ca 2+--independent phase, and CO2 insensitivity of the gca2 mutant. Proc. Natl. Acad. Sci. USA 103:7506-7511.
- Schroeder, J.I. (2006). Physiology: nitrate at the ion exchange. Nature 442:877-878.
- Hashimoto, M., Negi, J., Young, J., Israelsson, M., Schroeder, J.I., Iba, K. (2006). Arabidopsis HT1 kinase controls stomatal movements in response to CO 2. Nature Cell Biol. 8:391-397.
- Colcombet, J., Boisson-Dernier, A., Ros-Palau, R., Vera, C.E., Schroeder, J.I. (2005). Arabidopsis somatic embryogenesis receptor kinases 1 and 2 are essential for tapetum development and microspore maturation. Plant Cell 17:3350-3361.
- Leonhardt, N., Kwak, J. M., Robert, N., Waner, D., Leonhardt, G. & Schroeder, J. I. (2004). Microarray expression analyses of Arabidopsis guard cells and isolation of a recessive ABA hypersensitive Protein Phosphatase 2C mutant. Plant Cell 16:596-615.
- Gong, J.I., Waner, D.A., Horie, T., Li, S.L., Horie, R., Abid, K.B. and J.I. Schroeder (2004). Microarray-based rapid cloning of an ion accumulation deletion mutant in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 101:15404-15409.
- Kwak, J.M., I.C. Mori, Z-M. Pei, N. Leonhardt, M.A. Torres, J.L. Dangl, R.E. Bloom, S. Bodde, J.D.G. Jones and J.I. Schroeder. (2003). NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J. 22:2623-2633.
- Gong, J-M., D.A. Lee and J.I. Schroeder. (2003). Long-distance root-to-shoot transport of phytochelatins and cadmium in Arabidopsis. Proc. Natl. Acad. Sci. USA 100:10118-10123.
- Lahner, B, J. Gong, M. Mahmoudian, E.L. Smith, K.B. Abid, E.E. Rogers, M.L. Guerinot, J.F. Harper, J.M. Ward, L. McIntyre, J.I. Schroeder and D.E. Salt. (2003). Genomic scale profiling of nutrient and trace elements in Arabidopsis thaliana. Nature Biotechnology 21:1215-1221.
- Kwak, J.M., J-H. Moon, Y. Murata, K. Kuchitsu, N. Leonhardt, A. DeLong and J.I. Schroeder. (2002). Disruption of a guard cell-expressed protein phosphatase 2A regulatory subunit, RCN1, confers abscisic acid insensitivity in Arabidopsis. Plant Cell 14:2849-2861.
- Mäser, P., Y. Hosoo, S. Goshima, T. Horie, B. Eckelman, K. Yamada, K. Yoshida, E.P. Bakker, A. Shinmyo, Soiki, J.I. Schroeder, and N. Uozumi (2002). Glycine residues in potassium channel-like selectivity filters determine potassium selectivity in four-loop-per-subunit HKT transporters from plants. Proc. Nat. Acad. Sci. USA 99:6428-6433.
- Hugouvieux, V., J.M. Kwak and J.I. Schroeder. (2001). A mRNA cap binding protein, ABH1, modulates early abscisic acid signal transduction in Arabidopsis. Cell 106:477-487.
- Allen, G.J., S.P. Chu, C.L. Harrington, K. Schumacher, T. Hoffmann, Y.Y. Tang, E. Grill and J.I. Schroeder. (2001). A defined range of guard cell calcium oscillation parameters encodes stomatal movements. Nature 411:1053-1057.
- Schroeder, J.I., J.M. Kwak. and G.J. Allen. (2001). Guard cell abscisic acid signalling and engineering of drought hardiness in plants. Nature 410:327-330.
- Allen, G.J., Chu, S.P., Schumacher, K., Shimazaki, C.T., Vafeados, D., Kemper, A., Hawke, S.D., Tallman, G., Tsien, R.Y., Harper, J.F., Chory, J. and Schroeder, J.I. (2000). Alteration of Stimulus Specific Guard Cell Calcium Oscillations and Stomatal Closing in Arabidopsis det3 Mutant. Science 289:2338-2342.
- Pei, Z-M., Murata, Y., Benning, G., Thomine, S., Klüsener, B., Allen, G., Grill, E., and Schroeder, J.I. (2000). Calcium channels activated by hydrogen peroxide mediate abscisic acid signalling in guard cells. Nature 406:731-734.
- Clemens, S., E.J. Kim, D. Neumann and J.I. Schroeder. (1999). Tolerance to toxic metals by a gene family of phytochelatin synthases from plants and yeast. EMBO J. 18:3325-3333.
- Clemens, S., Antosiewicz, D.M., Ward, J.M., Schachtman, D.P. and Schroeder, J.I. (1998) The plant cDNA LCT1 mediates the uptake of calcium and cadmium in yeast. Proc. Natl. Acad. Sci. USA 95:12043-12048.
- Pei, Z-M., Ghassemian, M., Kwak, C.M., McCourt, P. and Schroeder, J.I. (1998) Role of farnesyltransferase in ABA regulation of guard cell anion channels and plant water loss. Science 282:287-290.
- Hildebrand, M., Volcani, B.E., Gassmann, W. and Schroeder, J.I. (1997) A gene family of silicon transporters. Nature 385:688-689.
- Pei, Z.M., Kuchitsu, K., Ward, J.M., Schwarz, M. and Schroeder, J.I. (1997) Differential abscisic acid regulation of guard cell slow anion channels in Arabidopsis wild-type and abi1 and abi2 mutants. Plant Cell 9:409-423.
- Pei, Z.M., Ward, J.M., Harper, J.F. and Schroeder, J.I. (1996) A novel chloride channel in Vicia faba guard cell vacuoles activated by the serine threonine kinase CDPK. EMBO J. 15:6564-6574.
- Rubio, F., Gassmann, W. and Schroeder, J.I. (1995). Sodium-driven potassium uptake by the plant potassium transporter HKT1 and mutations conferring salt tolerance. Science 270:1660-1663.
- Schachtman, D.P. and Schroeder, J.I. (1994). Structure and transport mechanism of a high-affinity K+ uptake transporter from higher plants. Nature 370:655-658.
Biography
Julian Schroeder did his PhD research at the Max Planck Institute for Biophysical Chemistry with Erwin Neher and was a Feodor von Lynen Humboldt Foundation postdoctoral fellow at UCLA School of Medicine. He received awards, including the Presidential Young Investigator Award (NSF), the ASPB Charles Albert Shull Award (1997), a DFG Heinz-Maier-Leibnitz Prize, the Blasker Award in Environmental Science, the ASPB Stephen Hales Prize (2021), an Alexander von Humboldt and Carl Friedrich von Siemens Research Prize (2022), is Churchill Overseas Fellow at Cambridge University (2019-) and with collaborators shared the Cozzarelli Prize from PNAS (2010) and a top 10 breakthrough of the year selected by Science (2009). He has served on several advisory boards, including Co-Director of the Food and Fuel for the 21st Century Center. He was von Humboldt Fellow at the TU Munich and the MPI for Biochemistry, visiting Professor at the ETH Zurich and is a member of the U.S. National Academy of Sciences, the German National Academy of Sciences - Leopoldina and Fellow of American Association for the Advancement of Science.

