Kimberly Cooper
Research
I can get lost in the exhibits and drawers of a natural history museum, and to me the skeletons are the best part. It’s fascinating that different species are so easily recognized by the isolated bones that are left behind when the natural processes of decomposition and fossilization are complete. One can’t help wonder how the dolphin turned an arm into a flipper, how the aye-aye got its creepy long finger, and how the horse evolved to run gracefully on a single toe.
Decades of research in the mouse and chick model systems as well as analyses of human birth defects have unlocked some of the mysteries of how the limb skeleton is patterned along three coordinate axes from the shoulder to finger tips, thumb to pinky finger, and back of the hand to palm. Yet neither human syndromes, nor the sledgehammer approach to knocking out genes in the mouse replicate the extraordinary range of adaptive morphologies observed in diverse species. This suggests a vast and untapped source of information about limb development and plasticity exists in the 400 million year genetic experiment that’s been continuing since the first “fishapod” flopped onto land.

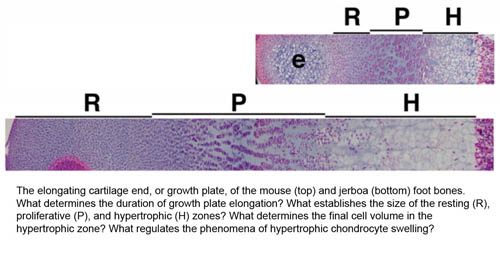
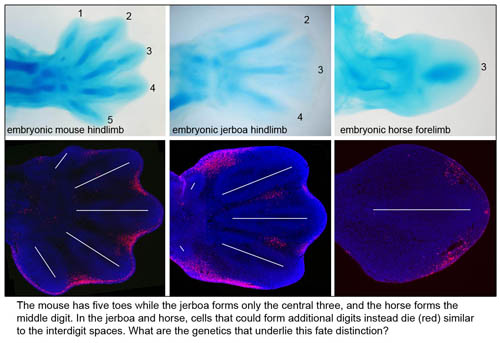
To gain a window into what evolution tells us about the developmental potential of the limb, we primarily focus on the three-toed jerboa, a small bipedal rodent that is closely related to the laboratory mouse and amenable to rearing in captivity. Compared to its quadrupedal relative, the jerboa has greatly elongated hindlimbs with three toes and metatarsals that fuse into a single bone. The unusual morphology of the jerboa skeleton allows us to ask what keeps one bone from growing into its adjacent neighbor? What mechanisms determine skeletal size and the relative proportions of individual elements? How are the correct numbers and positions of the digits established, and are the same mechanisms of digit loss redeployed in species that converge on similar morphologies (i.e. jerboas and horses, pigs and camels)? How can the forelimb and hindlimb evolve independently when both utilize the same developmental pathways? We apply classical embryology, cell biology, quantitative microscopy, high throughput sequence analysis, and mouse genetics in an integrative approach to understand these developmental mechanisms at the tissue and genome level.
Select Publications
- Villacís Núñez CN, Ray AP, Cooper KL, Moore TY (2022) Metatarsal fusion resisted bending as jerboas (Dipodidae) transitioned from quadrupedal to bipedal. Proceedings of the Royal Society B 289(1984): 20221322.
- Saxena A, Sharma V, Muthuirulan P, Neufeld SJ, Tran MP, Gutierrez HL, Erberich JM, Birmingham A, Cobb J, Capellini TD, Hiller M, Cooper KL (2022) Interspecies transcriptomics identify genes that underlie disproportionate foot growth in jerboas. Current Biology 32(2): 289-303.
- Weitzel AJ, Grunwald HA, Levina R, Gantz VM, Hedrick SM, Bier E, Cooper KL (2021) Meiotic Cas9 expression mediates gene conversion in the male and female mouse germline. PLoS Biology 19(12): e3001478.
- Saxena A and Cooper KL (2021) Diversification of the vertebrate limb: sequencing the events. Current Opinion in Genetics and Development, 69:42-47.
- McKenna KZ and Cooper KL (2021) A developmental perspective of homology and evolutionary novelty. Current Topics in Developmental Biology, 141:1-38.
- Gutierrez HL, Tsutsumi R, Moore TY, Cooper KL (2019) Convergent metatarsal fusion in jerboas and chickens is mediated by similarities and differences in the patterns of osteoblast and osteoclast activities. Evolution & Development. 21: 320–329. Editor’s Choice article for the issue.
- Tran MP, Tsutsumi R, Erberich JM, Chen KD, Flores MD, Cooper KL. (2019) Evolutionary loss of foot muscle during development with characteristics of atrophy and no evidence of cell death. eLife. 8: e50645. DOI: https://doi.org/10.7554/eLife.50645 Highlighted in eLife digest.
- Grunwald HA*, Gantz VM*, Poplawski G*, Xu XS, Bier E, Cooper KL. (2019) Super-Mendelian inheritance mediated by CRISPR/Cas9 in the female mouse germline. Nature. 566(7742): 105-109. Highlighted in Science News and in Nature News and Comment.
- Tstutsumi, R., Tran, M.P., and Cooper, K.L. (2017) Changing While Staying the Same: Preservation of Structural Continuity During Limb Evolution by Developmental Integration. Review. Integrative and Comparative Biology.
- Moore, T.Y., Cooper, K.L., Biewener, A.A., Vasudevan, R. (2017) Unpredictability of escape trajectory explains predator evasion ability and microhabitat preference of desert rodents. Nature Communications 8: 440.
- Saxena, A. and Cooper, K.L. (2017) The origins, scaling, and loss of tetrapod digits. Review. Proc. Biol. Sci. 372(1713):20150482.
- Saxena A and Cooper KL (2016) Fin to limb within our grasp. New and Views. Nature. 537: 176–177.
- Moore TY, Organ CL, Edwards SV, Biewener AA, Tabin CJ, Jenkins FA Jr, Cooper KL. (2015) Multiple Phylogenetically Distinct Events Shaped the Evolution of Limb Skeletal Morphologies Associated with Bipedalism in the Jerboas. Curr Biol. 25(21): 2785-2794.
- Cooper KL and Shapiro MD (2015) Preface to the special issue on evolution and morphological diversity. Dev Dyn. 244(10): 1181-1183.
- Cooper KL (2015) Self-organization in the limb: a Turing mechanism for digit development. Review. Curr Opin Genet Dev. 32: 92-97.
- Cook JA, Edwards SV, Lacey E, Guralnick RP, Soltis PS, Soltis DE, Welch CK, Bell KC, Galbreath KE, Himes C, Allen JM, Heath TA, Carnaval AC, Cooper KL, Liu M, Hanken J, Ickert-Bond S (2014) Aiming Up: Natural History Collections as Emerging Resources for Innovative Undergraduate Education in Biology. Bioscience. 64(8): 725-734.
- Cooper KL, Sears KE, Uygur A, Maier J, Baczkowski KS, Brosnahan M, Antczak D, Skidmore JA, Tabin CJ (2014) Patterning and post-patterning modes of evolutionary digit loss in mammals. Nature. 511(7507): 41-45.
- Cooper KL, Oh S, Sung Y, Dasari R, Kirschner MK, Tabin CJ (2013) Multiple phases of chondrocyte enlargement underlie differences in skeletal proportions. Nature 495: 375-378.
- Cooper KL (2011) The Lesser Egyptian Jerboa, Jaculus jaculus: A unique rodent model for evolution and development. Cold Spring Harbor Protocols (12): 1451-1456.
- Jordan B, Vercammen P, Cooper KL (2011) Husbandry and breeding of the Lesser Egyptian Jerboa, Jaculus jaculus. Cold Spring Harbor Protocols (12): 1457-1461.
- Cooper KL, Hu J, ten Berge D, Fernandez-Teran M, Ros M, Tabin CJ (2011) Initiation of proximal-distal patterning of the vertebrate limb by signals and growth. Science. 332(6033): 1083-1086.
Biography
Kimberly Cooper completed her PhD at the Fred Hutchinson Cancer Research Center in Seattle, WA. She then carried out her postdoctoral studies in the Department of Genetics at Harvard Medical School. She joined the Division of Biological Sciences faculty at U.C. San Diego in November 2013.

