Andrew Chisholm
Research
How the skin and nervous system maintain and repair themselves
The skin (epidermis) and the nervous system are each capable of regenerative responses to injury. Epidermal epithelia undergo robust wound healing responses in response to physical injury, related to the skin’s ability to maintain its long-term integrity. Neurons can also respond rapidly to damage and undergo regenerative regrowth leading to restoration of connectivity and function.
We study these processes in the nematode worm Caenorhabditis elegans, a tractable model for many aspects of skin and nervous system development and function. The C. elegans skin is a simple epithelium that encloses the entire animal during embryonic development and subsequently grows in size and complexity. The skin generates the cuticle, an apical extracellular matrix that mediates many skin functions and which serves as the animal’s exoskeleton. The nervous system contains only 302 neurons with simple stereotyped morphology. Neurons are closely associated with the epidermis, which has functions analogous to those of glial support cells.
Skin wound healing
The epidermis forms the first line of defense against environmental challenges including many microbial pathogens. In collaboration with Nathalie Pujol and Jonathan Ewbank (Marseille) we developed the C. elegans model for epidermal wound responses and repair and identified Death Associated Protein Kinase (DAPK) as a coordinate regulator of damage responses. We further discovered that DAPK acts via the epidermal microtubule cytoskeleton (Chuang et al., 2016, eLife). We have elucidated how the epidermis detects wounds, focusing on immediate damage response signals such as calcium, mitochondrial morphology, and reactive oxygen species (Xu and Chisholm, 2014, Dev. Cell). Current projects are exploring how the skin’s apical extracellular matrix (cuticle) responds to injury and repairs itself (see review by Zheng et al 2020, Faculty Reviews).
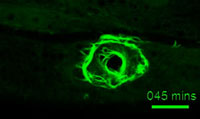
A ring of actin bundles forms after wounding (Suhong Xu)
Axon regeneration
The ability of axons to regrow after injury has been known for over a century and has been studied in a wide range of animals. However the molecular mechanisms of axon regeneration remain poorly understood owing in part to the lack of genetically tractable models. In collaboration with Yishi Jin (UCSD) we showed that C. elegans axons are capable of robust axon regeneration (Wu et al., 2007, PNAS). C. elegans axon regeneration requires the DLK-1 MAP kinase pathway, second messenger cascades, and axonal microtubule dynamics. We exploited the tractable genetics of C. elegans to test over 1500 genes for roles in axon regrowth and identified many genes with previously unknown roles as positive or negative regulators of regrowth (Chen et al., 2011, Neuron). The function of axon regeneration in the natural environment of C. elegans is not yet clear but may be related to the capacity of neurons to maintain their shape over the life of the animal. Current projects focus on factors such as DIP-2 (Noblett et al., 2019, J Cell Biol.) or CAR-1 (Tang et al 2020, Curr. Biol.) that regulate the shape of mature neurons.
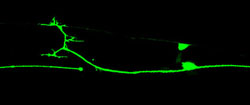
Axon regeneration after laser injury (Lizhen Chen)
Select Publications
Chisholm Lab selected publications:
Skin Wound Healing
- Xu, S., and Chisholm, A.D. 2011. A Gαq – Ca2+ signaling pathway promotes actin-mediated epidermal wound closure in C. elegans. Curr. Biol. 21: 1960-7. PMID: 25313960.
- Xu, S., and Chisholm, A.D. 2014. Skin wounding triggers a mitochondrial ROS burst that promotes wound repair. Dev. Cell. 31: 48-60. PMID: 25313960
- Chuang, M., Hsiao, T.I., Tong, A., Xu, S., Chisholm, A.D. 2016. DAPK interacts with Patronin and the microtubule cytoskeleton in C. elegans epidermal development and wound repair. eLife, Sep 23;5. pii: e15833. PMID: 27661253.
Reviews:
- Chisholm, A.D., and Xu, S. 2012. The C. elegans epidermis as a model skin II: differentiation and physiological roles. Wiley Interdiscip Rev Dev Biol. 1(6): 879-902. PMID: 23539358
- Li Zheng, S., Adams, J.G., and Chisholm A.D. 2020. Form and function of the apical extracellular matrix: new insights from Caenorhabditis elegans, Drosophila melanogaster, and the vertebrate inner ear. Faculty Reviews 2020 Dec 22; 9:27. doi: 10.12703/r/9-27 PMID: 33659959.
Axon Regeneration
- Chen, L., Wang, Z., Ghosh-Roy, A., Hubert, T., Yan, D., O’Rourke, S., Bowerman, B., Wu, Z., Jin, Y., Chisholm, A.D. 2011. Axon regeneration pathways identified by systematic genetic screening in C. elegans. Neuron 71: 1043-1057. PMID: 21943602
- Kim KW, Tang NH, Piggott CA, Andrusiak MG, Park S, Zhu M, Kurup N, Cherra SJ 3rd, Wu Z, Chisholm AD, Jin Y. 2018. Expanded genetic screening in Caenorhabditis elegans identifies new regulators and an inhibitory role for NAD+ in axon regeneration. eLife. pii: e39756. doi: 10.7554/eLife.39756. PMID: 3046142.
- Noblett N, Wu Z, Ding ZH, Park S, Roenspies T, Flibotte S, Chisholm AD, Jin Y, Colavita A. 2019. DIP-2 suppresses ectopic neurite sprouting and axonal regeneration in mature neurons. J Cell Biol. 218:125-133. doi: 10.1083/jcb.201804207. PMID: 30396999 eLife, Sep 23;5. pii: e15833. PMID: 27661253.
- Tang, N.H., Kim, K.W., Xu, S., Blazie, S.M., Yee, B.A., Yeo, G., Jin, Y., Chisholm, A.D. 2020. The mRNA decay factor CAR-1/LSM14 regulates axon regeneration via mitochondrial Ca dynamics. Curr. Biol. 30: 865-876. PMID: 31983639
Reviews:
- Chisholm, A.D., Hutter, H., Jin, Y., and Wadsworth, W.G. 2016. The genetics of axon guidance and axon regeneration in Caenorhabditis elegans. Genetics (WormBook). Nov;204(3):849-882. PMID: 28114100
For a complete list search for Andrew Chisholm on Google Scholar.
Biography
Andrew Chisholm received his PhD in 1989 from Cambridge University, working at the MRC Laboratory of Molecular Biology. He did postdoctoral research with H. Robert Horvitz at the Massachusetts Institute of Technology where he held a Lucille P Markey postdoctoral fellowship. He was on the faculty of the University of California Santa Cruz where he rose to the rank of Professor before moving to UC San Diego in 2007. From 2016-2019 he was Head of Cellular and Developmental Sciences at the Wellcome Trust in London. From 2019 he has served as Associate Dean of the Division of Biological Sciences.

