David Traver
Research
Utilizing zebrafish for new insights into the biology of hematopoietic stem cells
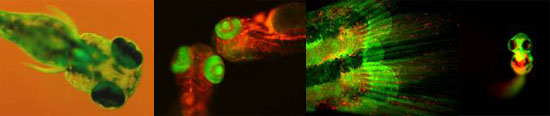
Hematopoietic stem cells (HSCs) exist as rare populations within blood-forming tissues that both self-renew and generate all blood cell lineages for life. HSCs have been isolated to relative purity in humans and mice, leading to a general understanding of their cellular and functional properties. Relatively little is known, however, regarding the genetic program underlying the embryonic formation and subsequent maintenance of HSC fate. The emergence of zebrafish as a model system has introduced unbiased genetic approaches, such as forward genetic screens and high-throughput expression screens, to the study of vertebrate blood cell formation. Fluorescent transgenesis technology combined with the external fertilization and transparency of zebrafish embryos allows early events in HSC specification, maintenance, and differentiation to be easily observed. Collectively, we are utilizing these advantages to image, fate map, and interrogate genetically the earliest embryonic blood-forming cells. This has helped address many long-standing questions in the development of the immune system, including the direct demonstration of HSC emergence from the transdifferentiation of aortic endothelium, and the demonstration of the importance of neighboring tissues in the instruction of HSC fate.
We are performing forward genetic and high-throughput expression screens to identify new gene functions in vertebrate stem cells. Novel genes that regulate HSC development, self-renewal, or lineage commitment will subsequently be examined for similar roles in mammalian systems. Despite decades of efforts, it is not yet possible to instruct HSC fate in vitro from pluripotent precursors. We hope that discovery of factors required to specify HSCs in the embryo will ultimately be useful in advancing this important goal of regenerative medicine.
Select Publications
- Sertori R, Liongue C, Basheer F, Lewis KL, Rasighaemi P, de Coninck D, Traver D, Ward AC. Conserved IL-2Ryc Signaling Mediates Lymphopoiesis in Zebrafish. J Immunol. (2016) 196:135-43.
- Butko E, Pouget C, Traver D. Complex regulation of HSC emergence by the Notch signaling pathway.Dev Biol. (2016) 409:129-38.
- Campbell C, Su T, Lau RP, Shah A, Laurie PC, Avalos B, Aggio J, Harris E, Traver D, Stachura DL. Zebrafish embryonic stromal trunk (ZEST) cells support hematopoietic stem and progenitor cell (HSPC) proliferation, survival, and differentiation. Exp Hematol. (2015) 43:1047-61.
- Kim BJ, Hancock BM, Bermudez A, Del Cid N, Reyes E, van Sorge NM, Lauth X, Smurthwaite CA, Hilton BJ, Stotland A, Banerjee A, Buchanan J, Wolkowicz R, Traver D, Doran KS. Bacterial induction of Snail1 contributes to blood-brain barrier disruption. J Clin Invest. (2015) 125:2473-83.
- Butko E, Distel M, Pouget C, Weijts B, Kobayashi I, Ng K, Mosimann C, Poulain FE, McPherson A, Ni CW, Stachura DL, Del Cid N, Espín-Palazón R, Lawson ND, Dorsky R, Clements WK, Traver D. Gata2b is a restricted early regulator of hemogenic endothelium in the zebrafish embryo. Development. (2015) 142:1050-61.
- Sahai-Hernandez P, Traver D. Intimacy of the niche: perivascular remodeling cuddles incoming HSCs.Cell Stem Cell. (2015) 16:109-10.
- Kim BJ, Hancock BM, Del Cid N, Bermudez A, Traver D, Doran KS. Streptococcus agalactiae infection in zebrafish larvae. Microb Pathog. (2015) 79:57-60.
- Kim AD, Stachura DL, Traver D. Cell signaling pathways involved in hematopoietic stem cell specification. Exp Cell Res. (2014) 329:227-33. Review.
- Pouget C, Peterkin T, Simões FC, Lee Y, Traver D, Patient R. FGF signaling restricts haematopoietic stem cell specification via modulation of the BMP pathway. Nature Commun. (2014) 5:5588.
- Lee Y, Manegold JE, Kim AD, Pouget C, Stachura DL, Clements WK, Traver D. FGF signaling specifies haematopoietic stem cells through its regulation of somitic Notch signaling. Nature Commun. (2014) 5:5583.
- Espín-Palazón R, Stachura DL, Campbell CA, García-Moreno D, Del Cid N, Kim AD, Candel S, Meseguer J, Mulero V, Traver D. Proinflammatory signaling regulates hematopoietic stem cell emergence. Cell. (2014) 159:1070-85.
- Kim AD, Melick CH, Clements WK, Stachura DL, Distel M, Panáková D, MacRae C, Mork LA, Crump JG, Traver D. Discrete Notch signaling requirements in the specification of hematopoietic stem cells .EMBO J. (2014) 33:2363-73.
- Ouyang K, Leandro Gomez-Amaro R, Stachura DL, Tang H, Peng X, Fang X, Traver D, Evans SM, Chen J.Loss of IP3R-dependent Ca2+ signaling in thymocytes leads to aberrant development and acute lymphoblastic leukemia. Nature Commun. (2014) 5:4814.
- Stachura DL, Traver D. Hematopoietic ontogeny in the axolotl. Blood. (2014) 124:1204-6.
- Kobayashi I, Kobayashi-Sun J, Kim AD, Pouget C, Fujita N, Suda T, Traver D. Jam1a-Jam2a interactions regulate haematopoietic stem cell fate through Notch signaling. Nature. (2014) 512:319-23.
- Wittamer, V., J.Y. Bertrand, P.W. Gutschow, and D. Traver. 2011. Characterization of the mononuclear phagocyte system in zebrafish. Blood, Epub March 15.
- Clements, W.C., A.D. Kim, K.G. Ong, J.C. Moore, N.D. Lawson, and D. Traver. 2011. A somitic Wnt16/Notch pathway specifies haematopoietic stem cells. Nature, 474: 220-224.
- Clements, W.C., and D. Traver. 2013. Signaling pathways that control vertebrate haematopoietic stem cell specification. Nature Reviews Immunology, 13: 336-348.
- Bertrand, J.Y., N.C. Chi, B. Santoso, S. Teng, D.Y.R. Stainier, and D. Traver. 2010. Hematopoietic stem cells derive directly from aortic endothelium during development. Nature, 464: 108-111.
Biography
David Traver received his Ph.D. in immunology from Stanford University where he received the McDevitt Prize for best thesis. He then completed postdoctoral studies at Harvard University as a fellow of the Irvington Institute. Since joining UCSD in 2004, he has received a Young Investigator Award from the Beckman Foundation, a Scholar Award from the American Society of Hematology and a New Faculty Award from the California Institute for Regenerative Medicine.

