Ella Tour
Research
Education
Critical thinking skills, such as the ability to analyze data, solve problems, and develop new ideas, are crucial for success in the modern workforce. Although the majority of biology educators consider critical and creative scientific thinking some of the most important skills to be acquired as a result of science education, few biology faculty report spending enough time practicing these skills in their classrooms (Coil et al., 2010). One of my primary education research goals is to increase our understanding of how to effectively teach critical thinking skills in biology and how objectively test whether students are learning these skills. In my classes, I strive to provide my students with ample opportunities to practice data analysis and experimental design. Currently, my collaborators and I are investigating the effectiveness of in-depth analysis of primary research papers and designing experiments that follow up on these papers, on students' critical thinking skills. An additional area of education research is developing methods to improve the quality of scientific writing and prevent plagiarism in Biology labs. This research is performed in collaboration with Tricia Bertram Gallant (UCSD Office of Academic Integrity), Julian Parris (Psychology), Goran Bozinovic (Biology), and Madeline Picciotto (UCSD Writing Center).
Biological
In collaboration with the members of Bill McGinnis's lab here at UCSD, I study the role of chromatin structure and long non-coding RNA's in the regulation of gene expression in developing embryos of the fruit fly Drosophila melanogaster. Early embryonic development is a precisely orchestrated process in which genes that control the formation of the various body parts have to be expressed at the right time and the right place. One such master control gene, Ultrabithorax ( Ubx) is responsible for the correct development of parts of fly's thorax and abdomen. The expression of Ubx is regulated by multiple enhancers that are located tens of thousands base pairs away from the Ubx promoter and its transcription start site. Using high-resolution fluorescent in situ hybridization (FISH), we study the conformational changes involving Ubx promoter and its enhancers that occur in silent and transcriptionally active states . We are also investigating a potential role of an antisense transcript that originates in one of the introns of Ubx in the regulation of this gene.
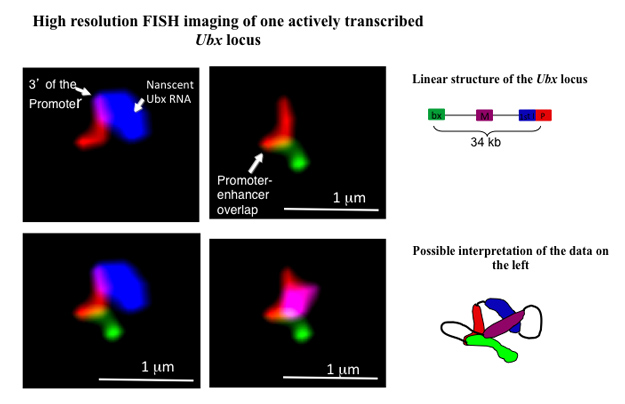
This image shows one actively transcribed Ubx gene in 3 hours-old Drosophila embryo. P (red) – Ubx promoter, mRNA (blue) – nascent Ubx mRNA originating from the 1st intron of this gene, bx (green) – bx enhancer, M (purple) – a region located midway between the promoter and bx enhancer. bx enhancer, located 34 kbp downstream of Ubx promoter, appears to be in contact with this promoter.
Courses
BGGN 211: Recent Advances and Experimental Approaches in Modern Biology
This is a graduate-level course open to the Master's students enrolled in the Contiguous BS/MS program in Biology. This 4 units course aims to provide rigorous training in critical analysis of scientific papers, data interpretation, scientific writing, and experimental design. Through written critiques and in-class discussions, students analyze the experimental data presented in several recent papers from different fields of biology. Working in small groups, they design a controlled experiment that follows up on the research described in the papers, and evaluate experimental proposals by other groups. Students also write a mini-review on the topic of their research that can serve as the basis to the Introduction section of their Master's thesis.
BIMM 101: Recombinant DNA Techniques Labs
In this upper-division Molecular Biology lab course, students learn about and perform the basic approaches to working with DNA: gel electrophoresis, restriction digests, ligations, transformations, extraction of genomic DNA and mRNA, and PCR. Experiments include isolating the lux operon, a group of genes that can produce light, inducing and quantifying RNA interference in C. elegans, and characterizing San Diego biodiversity using mitochondrial DNA. Some of the main goals of this class include developing skills of data analysis, effective data presentation, and scientific writing.
BIMM 110: Molecular Basis of Human Disease
This upper-division lecture course presents the current understanding of the molecular underpinning and the available treatments of several representative human diseases, such as Hemophilia, Cancer, Diabetes, Alzheimer's disease, and others. Since our understanding of human diseases develops at an extremely rapid pace, most of the material presented in this course is taken from recent reviews and research articles. To increase material retention and students' engagement in this large class, I incorporate active-learning teaching methods such as Peer-Instruction (Crouch and Mazur, 2001), combined with the use of iClickers and group work, where students analyze data and design experiments. In discussion sections that accompany the lectures, students practice analyzing research papers that investigate diseases using human populations and model organisms, such as mice, fruit flies, and worms.
Select Publications
- Bertram Gallant T , Picciotto M, Bozinovic G, and Tour E. (2019) Plagiarism or Not? Investigation of Turnitin®‐Detected Similarity Hits in Biology Laboratory Reports. Biochem Mol Biol Educ. doi:10.1002/bmb.21236
- Lie R, Abdullah C, He W, Tour E. (2016) Perceived Challenges in Primary Literature in a Master’s Class: Effects of Experience and Instruction. CBE Life Science Education Volume 15: ar77. pmid:27909027
- Abdullah C, Parris J, Lie, R, Guzdar A, and Tour E (2015) Critical analysis of primary literature in a Master’s-level class: effects on self-efficacy and science process skills. CBE-Life Sciences Education, Volume 14, 1-13.
- Brydges S, Chilukuri L, Cook G, Feeley M, Herbst M, Tour E, and Van Den Einde L* (2012). Developing a Faculty Learning Community at a Large Research University: Interdisciplinary Connections and Competencies in the Scholarship of Teaching and Learning. Currents in Teaching and Learning Spring 2013. Volume 6.
- Tour E and McGinnis W. (2006) Gap peptides: A new way to control embryonic patterning? Cell, 126(3): 448-9.
- Tour E, Hittinger CT, and McGinnis W. (2005) Evolutionarily conserved domains required for activation and repression functions of the Drosophila Ubx protein. Development, 132: 5271-5281.
- Tour E and McGinnis W. (2003) Hox Genes and Evolution of Body Plan. In: Encyclopedia of the Human Genome. Macmillan Publishers Ltd, Nature Publishing Group.
- Tour E, Pillemer G, Gruenbaum Y, and Fainsod A. (2002) Gbx2 interacts with Otx2 and patterns the anterior-posterior axis during gastrulation in Xenopus. Mech Dev. 112(1-2): 141-51.
- Tour E, Pillemer G, Gruenbaum Y, and Fainsod A. (2002) Otx2 can activate the isthmic organizer genetic network in the Xenopus embryo. Mech Dev. 110(1-2): 3-13.
- Tour E, Pillemer G, Gruenbaum Y, and Fainsod A. (2001) The two Xenopus Gbx2 genes exhibit similar, but not identical expression patterns and can affect head formation. FEBS Lett. 507(2):205-9.
* Authors are listed in alphabetical order
Biography
Ella Tour received her Ph.D. in Genetics at the Hebrew University of Jerusalem, where she studied the molecular mechanisms of brain development in the frog Xenopus laevis. She conducted her post-doctoral training at UCSD, investigating the early development of the fruit fly embryos.

