Matthew Banghart
Research
How do cognitive processes such as expectation and attentional focus shape sensory perception? From where in the brain do predictions emerge, and how are they relayed to sensory circuits to impact perception? What neurotransmitters are involved and what is the role of synaptic plasticity? My lab aims to answer these questions in the context of pain perception (e.g. placebo analgesia), such that mechanistic answers will not only provide fundamental insights, but may also point the way to new therapies. Although our approach relies largely on contemporary optogenetic and chemogenetic methods, brain slice electrophysiology, in vivo imaging, and rodent behavioral analysis, it is distinguished by in-house development of novel photochemical tools that are customized to the questions at hand.
Research in the lab is centered around understanding how cognitive processes influence pain perception by recruiting endogenous opioid signaling throughout the brain. Although functional imaging studies in humans implicate multiple brain structures in expectation-driven pain relief, due to a lack of mechanistic studies in model organisms, current models of top-down pain modulation are based almost entirely on correlative data. Currently, our primary goals are to delineate the neural pathways that support placebo analgesia, decipher their computational roles, and uncover the function of opioid neuropeptide signaling in the underlying circuits. Ultimately, we hope to leverage this information to develop novel behavioral and pharmacological therapies for pain.
To address the potential roles of endogenous opioid signaling (e.g. encoding prediction precision), we are optimizing our light-activated opioids for in vivo applications. We are currently developing small molecule agonists and antagonists that can be administered systemically and activated through implanted fibers during placebo trials to test the impact of local, graded, and temporally-delimited opioid signaling on the placebo effect. These efforts will be complemented by implementation of emerging genetically-encoded sensors of endogenous opioid neuropeptides. In parallel, we are pursuing a related effort that is uncovering new signaling pathways that govern neuropeptide release. These studies focus on the brain areas implicated in opioid-dependent cognitive pain modulation, which promises to reveal the complex interplay of neuromodulatory signaling that shapes sensory perception.
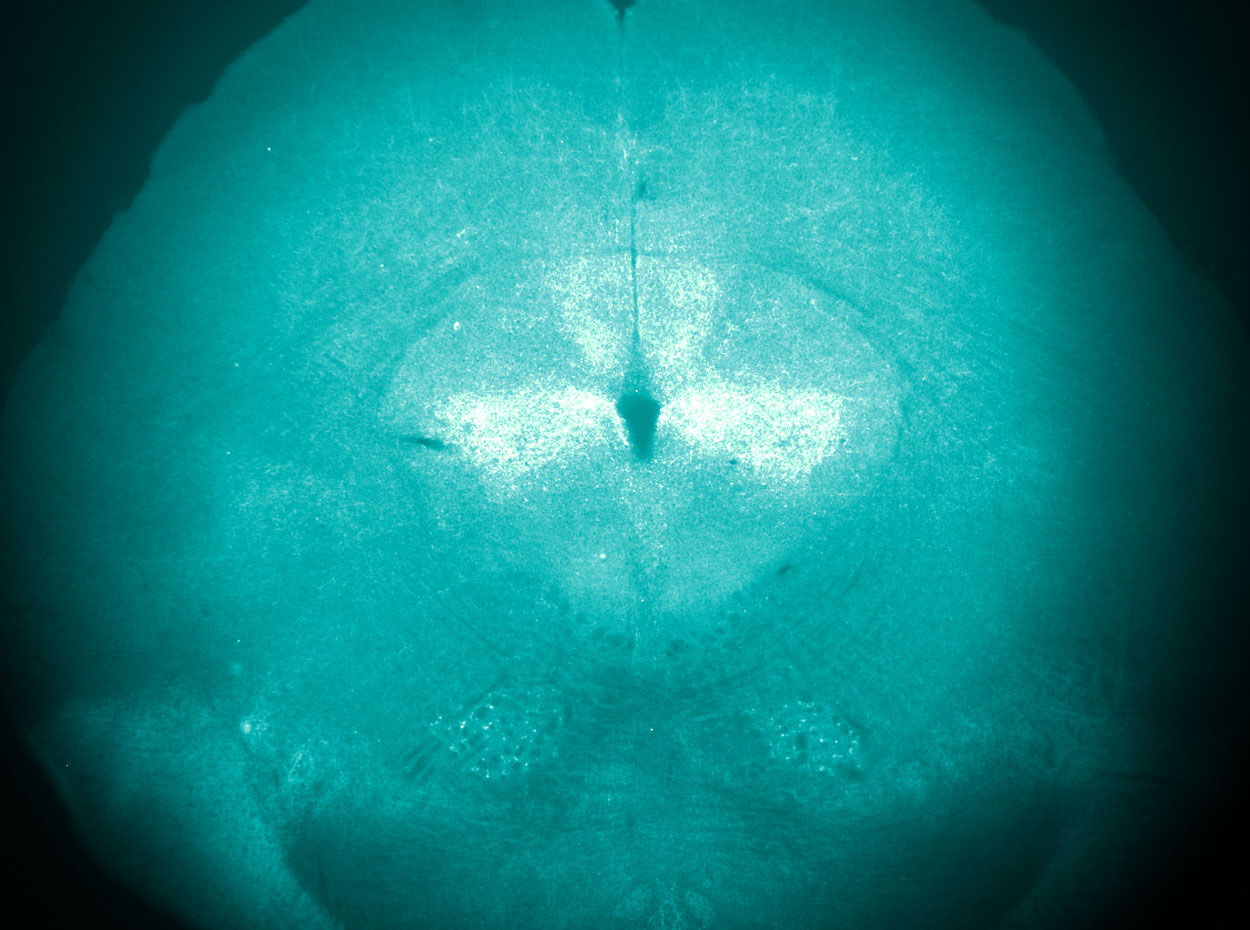
Anti-nociceptive neuropeptide expression in the periaqueductal gray, a central node of the descending pain modulatory pathway.
Select Publications
- Banghart MR, He XJ, Sabatini BL. A Caged Enkephalin Optimized for Simultaneously Probing Mu and Delta Opioid Receptors. ACS Chem Neurosci. 2017 Dec 27. PMID: 29266926
- Otopalik AG, Sutton AC, Banghart M, Marder E. When complex neuronal structures may not matter. Elife. 2017 Feb 6;6. pii: e23508. PMID: 28165322
- Banghart MR, Neufeld SQ, Mulder N, Sabatini BL. Enkephalin disinhibits mu-receptor rich striatal patches via delta opioid receptors. Neuron (2015), 88(6), 1227-39. PMID 22671460
- Olson J, Banghart MR, Sabatini BL, Ellis-Davies GCR. Spectral evolution of a photochemical protecting group for orthogonal two-color uncaging with visible light. Journal of the American Chemical Society (2013), 135(42), 15948-54. PMID 24117060
- Banghart MR, Williams JT, Shah RC, Lavis LD, Sabatini BL. Caged naloxone reveals opioid signaling deactivation kinetics. Molecular Pharmacology (2013), 84(5), 687-95. PMID 23960100
- Banghart MR, Sabatini BL. Photoactivatable neuropeptides for spatiotemporally precise delivery of opioids in neural tissue. Neuron (2012), 73, 249-59. PMID 22284180
- Tochitsky I, Banghart MR, Mourot A, Yao JZ, Gaub B, Kramer RH, Trauner D. Optical control of genetically engineered neuronal nicotinic acetylcholine receptors. Nature Chemistry (2012), 4(2), 105-11. PMID 22270644
- Dvir T, Banghart MR, Timko BP, Langer R, Kohane DS. Photo-targeted nanoparticles. Nano Letters (2010), 10 (1), 250-254. PMID 19904979
- Banghart MR, Mourot A, Fortin DL, Yao JZ, Kramer RH, Trauner D. Photochromic blockers of voltage-gated potassium channels. Angewandte Chemie International Edition (2009), 48 (48), 9097-9101. PMID 19882609
- Fortin DL, Banghart MR, Dunn TW, Borges K, Wagenaar DA, Gaudry Q, Karakossian MH, Otis TS, Kristan WB, Trauner D, Kramer RH. Photochemical control of endogenous ion channels and cellular excitability. Nature Methods (2008), 5(4), 331-338. PMID 18311146
- Banghart MR, Volgraf M, Trauner D. Engineering light gated ion channels. Biochemistry (2006), 45(51), 15129-15141. PMID 17176035
- Banghart M, Borges K, Isacoff E, Trauner D, Kramer RH. Light-activated ion channels for remote control of neuronal firing. Nature Neuroscience (2004), 7(12), 1381-1386. PMID 15558062
- Miller AK, Banghart MR, Beaudry CM, Suh JM, Trauner D. Development of novel Lewis acid catalyzed cycloisomerizations: synthesis of bicyclo[3.1.0]hexenes and cyclopentenones. Tetrahedron (2003), 59(45), 8919-8930.
Biography
Matt Banghart obtained his Ph.D. in Chemistry from UC Berkeley and pursued postdoctoral training in the Department of Neurobiology at Harvard Medical School with Bernardo Sabatini. Matt was a Helen Hay Whitney Postdoctoral Fellow and the recipient of a K99/R00 Pathway to Independence Award from NIDA. He is currently establishing his independent research laboratory at UCSD.

